Large Epilepsy Genetics Study Compares Severe and Less-severe Epilepsies in 17,606 Individuals
July 22, 2019
Sequencing-based studies have identified novel risk genes associated with severe epilepsies and revealed an excess of rare deleterious variation in less-severe forms of epilepsy. To identify the shared and distinct ultra-rare genetic risk factors for different types of epilepsies, we performed a whole-exome sequencing (WES) analysis of 9,170 epilepsy-affected individuals and 8,436 controls of European ancestry.
The research team focused on three phenotypic groups: severe developmental and epileptic encephalopathies (DEEs), genetic generalized epilepsy (GGE), and non-acquired focal epilepsy (NAFE).
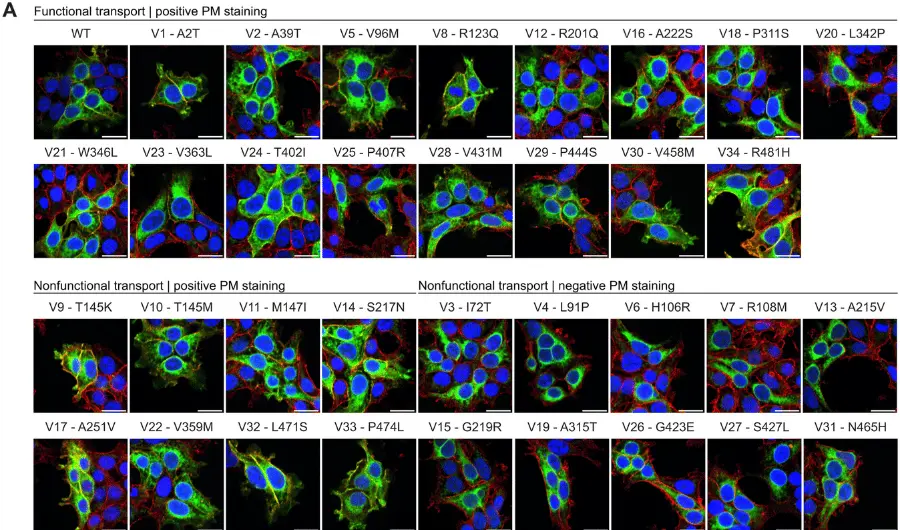
The team observed that compared to controls, individuals with any type of epilepsy carried an excess of ultra-rare, deleterious variants in constrained genes and in genes previously associated with epilepsy; they saw the strongest enrichment in individuals with severe developmental and epileptic encephalopathies and the least strong in individuals with non-acquired focal epilepsy. Moreover, the research team found that inhibitory GABA A receptor genes were enriched for missense variants across all three classes of epilepsy, whereas no enrichment was seen in excitatory receptor genes. The larger gene groups for the GABAergic pathway or cation channels also showed a significant mutational burden in DEEs and GGE. Although no single gene surpassed exome-wide significance among individuals with GGE or NAFE, highly constrained genes and genes encoding ion channels were among the lead associations; such genes included CACNA1G,EEF1A2, and GABRG2 for GGE and LGI1, TRIM3, andGABRG2 for NAFE.
The research team’s study, the largest epilepsy whole-exome sequencing study to date, confirms a convergence in the genetics of severe and less-severe epilepsies associated with ultra-rare coding variation, and it highlights a ubiquitous role for GABAergic inhibition in epilepsy etiology.